Phylogenetic Analysis of Terminal Restriction Fragments (tRFs)
[under construction]
TReFID is a tool for analyzing terminal restriction fragments of multiple restriction enzyme digests. It allows to create "species lists" on the basis of tRFLP data from complex microbial communities. An impression on how it works is given by figures 1 through 5, below. [AEM paper on TReFID] [Download TReFID]
Currently databases for three genes are available (included in the download packages, see below):
-
bacterial small subunit rRNA gene (SSU = 16S rDNA)
-
dinitrogenase reductase gene (nifH)
-
nitrous oxide reductase gene (nosZ)
| TReFID software package version 1.0 | first release; database versions A (9.3 MB) |
| TReFID software package version 1.2.1 | improved software; expanded databases, versions B (14.5 MB) |
Note: The TReFID software has no web interface; all required components - including databases for the three genes - are included in these packages.
In case of nifH and nosZ all sequences published in GenBank (Jan 2004 and Feb 2005, respectively) were tested for TReFID. Only sequences containing the binding site for the fluorochrome-labelled PCR primers used in tRFLP analyses were included in the databases. These were the primers nifH-F (aaa ggY ggW atc ggY aaR tcc acc ac), binding near the 5' end of nifH, and nosZ-R (tcc atg tgc agN gcR tgg cag aa), binding close to the 3' terminus of nosZ. Because the nifH and nosZ genes have no motifs comparably highly conserved as some regions in the SSU gene, and because of the limited sequence information, primers targetting these genes are "less universal" than established SSU primers like 8F, 27F, or 63F. As a consequence, the primers nifH-F and nosZ-R allow to exploit only about half the available sequence information for the respective gene. However, this proportion would not have been larger using alternative primers. Concerning the SSU gene, about 40,000 out of 150,000 GenBank sequences published in Feb 2005 were tested. Of these, about 21,000 sequences contained in the TReFID SSU database (version B) included the binding site for 63F, the primer used for tRFLP. Database overview
| number of entries | date of last change | |
| SSU / 16S rDNA (version B) | 22,145 | 10-APR-2005 |
| SSU / 16S rDNA (version A) original version | 17,462 | 25-AUG-2004 |
| nifH | 1,318 | 10-APR-2005 |
| nosZ | 607 | 10-APR-2005 |
Note: all three databases are included in the ZIP packages above and need not be downloaded separately. This is for future updates or new databases. New databases can be added by copying the unzipped .TReFID.org files into the TReFID\source subdirectory. Existing databases can be altered and extended with user-specific data in any text editor.
The figures below are taken from the original publication on TReFID (C Rösch & H Bothe. 2005. Appl. Environ. Microbiol. 71(4) 2026-2035). To obtain high quality images of these figures click on the respective image. For figure legends and detailed background information follow the links.
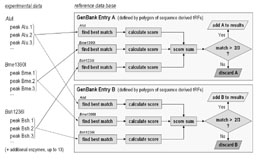 |
Figure 1: Flow chart describing the steps in the identification procedure of the TReFID program. |
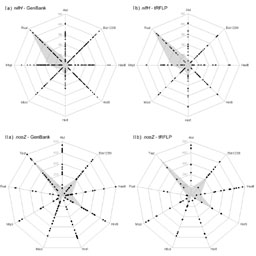 |
Figure 2: Graphical representation of the tRFs obtained for nifH of N2-fixing and nosZ of denitrifying microorganisms. |
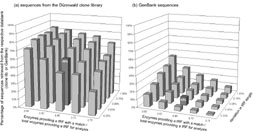 |
Figure 3: Identification of polygons representing organisms in the Dünnwald soil in two libraries. |
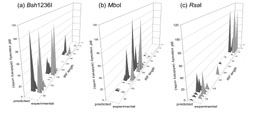 |
Figure 4: Comparison in the SSU tRF profiles obtained experimentally with DNA from the Dünnwald soil and predicted from the sequence information in the TReFID results list. |
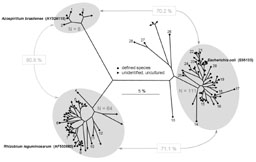 |
Figure 5: The occurrence of SSU tRF polygons related to those from either Azospirillum brasilense, Escherichia coli or Rhizobium leguminosarum. |
External links:
-
Angela D. Kent, Dan J. Smith, Barbara J. Benson & Eric W. Triplett (2003). Web-Based Phylogenetic Assignment Tool for Analysis of Terminal Restriction Fragment Length Polymorphism Profiles of Microbial Communities. Applied and Environmental Microbiology 69(11): 6768-6776. [link] [PAT site]
-
Christopher L. Kitts (2001). Terminal restriction fragment patterns: a tool for comparing microbial communities and assessing community dynamics. Current Issues in Intestinal Microbiology 2(1): 17-25. [abstract]
-
Wen-Tso Liu, Terence L. Marsh, Hans Cheng & Larry J. Forney (1997). Characterization of microbial diversity by determining terminal restriction fragment length polymorphisms of genes encoding 16S rRNA. Applied and Environmental Microbiology 63(11): 4516-4522. [link]
-
A. Mark Osborn
, Edward R. B. Moore & Kenneth N. Timmis (2000). An evaluation of terminal-restriction fragment length polymorphism (T-RFLP) analysis for the study of microbial community structure and dynamics. Environmental Microbiology 2(1): 39-50 [link]
-
Terence L. Marsh
, Paul Saxman, James Cole & James Tiedje (2000). Terminal Restriction Fragment Length Polymorphism Analysis Program, a Web-Based Research Tool for Microbial Community Analysis. Applied and Environmental Microbiology 66(8): 3616-3620. [link]
-
Microbial Community Analysis (MiCA) [link]
-
Ribosomal Database Project (RDP) [link]
-
Bellerophon Chimera Detection Tool [link]
-
Phylo_Win (Molecular Phylogenetic Inference by NJ, ML or Maximum Parsimony) [link]
-
BioEdit Biological Sequence Alignment Editor [link]
TReFID was written in PureBasic:

Thanks to all the helpful people on the PB forums!
TReFID arose from a project funded by:


Contact:
webmaster@trefid.net
(last update: 10-APR-2005)

